 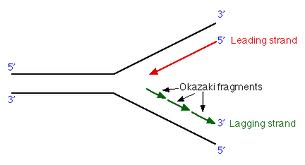
Hanoian, Philip Gannavaram, Swathi Benkovic, Stephen J. RNA primer–primase complexes serve as the signal for polymerase recycling and Okazaki fragment initiation in T4 phage DNA replication The similarities to both bacterial and eukaryotic systems and evolutionary implications of archaeal Okazaki fragment maturation are discussed. This flap structure was cleaved by flap endonuclease 1 (Fen1) and the resultant nick was ligated by DNA ligase to form a mature lagging strand. Instead, Family B DNA polymerase (polB) was observed to rapidly fill the gaps left by polD and displaces the downstream Okazaki fragment to create a flap structure. It is shown here, however, that it stops before the previous Okazaki fragments, failing to rapidly process them. DNA polymerase D (polD) was proposed to function as the replicative polymerase in Thermococcus replicating both the leading and the lagging strands. A dual color fluorescence assay was developed to monitor reaction substrates, intermediates, and products. To address the components required for the process in Thermococcus, Okazaki fragment maturation was reconstituted in vitro using purified proteins from Thermococcus species 9°N or cell extracts. Efficient maturation prevents repeat sequence expansions, small duplications, and generation of double-stranded DNA breaks. Greenough, Lucia Kelman, Zvi Gardner, Andrew F.ĭuring replication, Okazaki fragment maturation is a fundamental process that joins discontinuously synthesized DNA fragments into a contiguous lagging strand. 
The Roles of Family B and D DNA Polymerases in Thermococcus Species 9°N Okazaki Fragment Maturation* The RNA trimer may, therefore, lock the complete fragment in an A-type conformation. Although the base-pair geometry, particularly in the central TATA part, is distorted, there is no evidence for a transition from the A- to the B-type conformation at the junction between RNA.DNA hybrid and DNA duplex. 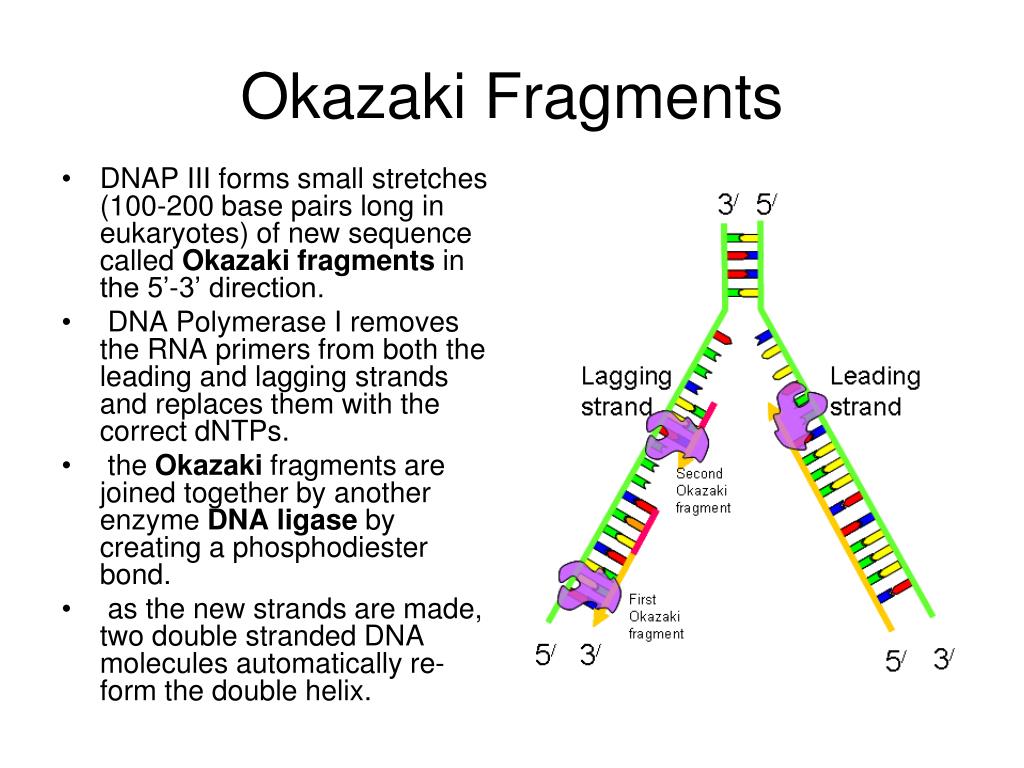
The fragment adopts an overall A-type conformation with 11 residues per turn. The DNA oligonucleotide d(GGGTATACGC) and the chimeric RNA-DNA oligonucleotide r(GCG)d(TATACCC) were combined to form a synthetic Okazaki fragment and its three-dimensional structure was determined by x-ray crystallography. They are composed of the growing DNA strand primed by RNA and the template strand. In DNA replication, Okazaki fragments are formed as double-stranded intermediates during synthesis of the lagging strand. This method combines PBM and primase activity assay and is designated as high-throughput primase profiling (HTPP), and it allows characterization of specific sequence recognition by DNA primase in unprecedented time and scalability.Crystal structure of an Okazaki fragment at 2-A resolutionĮgli, M. Using a high-throughput microarray-based binding assay and consecutive biochemical analyses, it has been shown that 1) the specific binding context (flanking sequences of the recognition site) influences the binding strength of the DNA primase to its template DNA, and 2) stronger binding of primase to the DNA yields longer RNA primers, indicating higher processivity of the enzyme. Conventional biochemical tools that are used to determine the DNA recognition sequence of DNA primase provide only limited information. It is a pivotal step in the formation of Okazaki fragments. The binding of prokaryotic DnaG-like primases to DNA occurs at a specific trinucleotide recognition sequence. DNA primase synthesizes short RNA primers that initiate DNA synthesis of Okazaki fragments on the lagging strand by DNA polymerase during DNA replication. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
